Mice
Male wild-type mice were bred in house and weaned at 3 weeks or obtained from Jackson Laboratories (stock no. 000664). β-arr2-KO mice (stock no. 011130) were obtained from Jackson Laboratories, bred in house and weaned at 3 weeks of age. All mice were inbred to the C57BL/6J congenic ‘wild-type’ strain (as opposed to outbred ‘true wilds’, which were not used in this study). Congenic strains are generated by backcrossing for a minimum of 10 generations, a standard that is derived from the congenic interval, and the theoretical estimate that by the 10th generation, 99.99% of the congenic strain background will be from the recipient inbred69. Although the β-arr2-KO mouse (Jackson Laboratories stock no. 011130), was originally derived on the 129X1/SvJ background70, it was backcrossed to the C57BL/6J congenic strain at Jackson Laboratories (https://www.jax.org/strain/011130). All mice were maintained on a 12 h:12 h natural light:dark cycle, starting at 07:30 with food and water provided ad libitum. All behavioural experiments were conducted during the same circadian period (07:30–19:30) in a dedicated, sound- and odour-controlled behavioural testing room, which is separated from the vivarium, and no other experiments were conducted simultaneously in the same room. Sample size was estimated based on previous work and published literature. Experimenters were blind to the condition when subjective criteria were used as a component of data analysis, and control and test conditions were interleaved. Mice were randomly assigned to experimental and control groups. All procedures complied with the animal care standards set forth by the National Institutes of Health and were in accordance with protocols approved by the Johns Hopkins University Animal Care and Use Committee.
sCPP assay
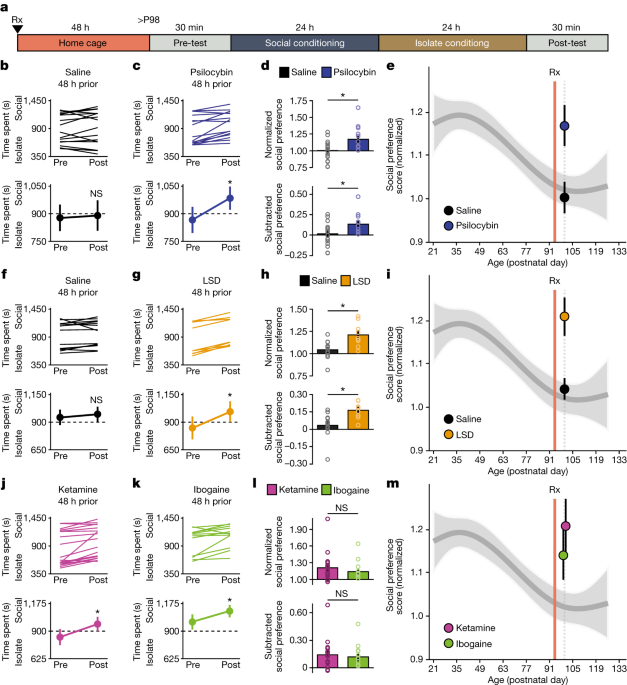
The protocol for sCPP was adapted from previously published work11. Mice were socially housed (3–5 males) in a cage containing corncob bedding (Anderson Cob, 0.25 inch cob, Animal Specialties and Provisions) until the pre-determined age for sCPP testing. Each mouse was used for only one behavioural time point. At the pre-determined age, mice were placed in an open field activity chamber (ENV-510, Med Associates) equipped with infrared beams and a software interface (Activity Monitor, Med Associates) to monitor the position of the mouse. The apparatus was partitioned into two equally sized zones using a clear Plexiglas wall, with a 5 cm diameter circular hole at the base; each zone contained one type of novel bedding (Alpha-Dri, Animal Specialties and Provisions or Kaytee Soft Granule, Petco). The amount of time spent freely exploring each zone was recorded during 30-min test sessions. For example, a score of 900 means that the mouse spent exactly 50% of its time on each of the two beddings, whereas a score of 1,800 means that it spent the full 30 min in the bedding that would be subsequently assigned as the social conditioning cue, and no time in the bedding that would be assigned as the isolation conditioning cue. After an initial pre-conditioning trial to establish baseline preference for the two sets of bedding cues, mice were assigned to receive social conditioning (with cage mates) for 24 h on one type of bedding, followed by 24 h of isolation conditioning (without cage mates) on the other bedding cue. To assure unbiased design, chamber assignments were counterbalanced for side and bedding cues. Immediately after the isolation conditioning, a 30-min post-conditioning trial was conducted to establish preference for the two conditioned cues. CPP is a learned association between a condition (for example, social) and a cue (bedding). It does not require scent from the other mice, as the bedding itself serves as the cue. Exclusion criteria for this behaviour are strictly defined as a pre-conditioning preference score of >1.5 or <0.5. Mice are never excluded based on the quality of their social interactions. Pre-conditioning versus post-conditioning social preference scores were considered significant if paired Student’s t-test P values were less than 0.05. Comparisons between experimental conditions were made using both normalized social preference scores (time spent in social zone post-treatment divided by pre-treatment) and subtracted social preference scores (time spent in social zone post minus pre); these were considered significant if unpaired Student’s t-test P values were <0.05. All experiments were performed during the mouse rest period (light cycle), since pilot experiments revealed that sCPP is most robust if assayed during this period. Prior to i.p. drug treatment experiments (MDMA, LSD, psilocybin, ketamine or ibogaine hydrochloride), mice were habituated to the injection procedure with daily saline i.p. injections in the home cage. Pharmacological delivery schedules were counterbalanced for type of drug. Unless otherwise stated (Fig. 2 and Extended Data Fig. 5), for pretreatment, experiments mice were tested 48 h after the injection to allow for complete clearance of the drug. For the experiment testing involvement of the 5-HT 2A R, the 5-HT 2A R antagonist ketanserin was administered i.p. 30 min prior to the injection of the drug tested.
Electrophysiology
Subjects received an i.p. injection of either LSD (1 µg kg−1), ketamine (3 mg kg−1), psilocybin (0.3 mg kg−1), MDMA (10 mg kg−1), ibogaine (40 mg kg−1) or saline. Forty-eight hours after drug treatment, either parasagittal slices containing the NAc core (250 µm thick) or coronal slices containing the PL/IL region of the mPFC (250 µm thick) were prepared from C57BL/6 mice using standard procedures. In brief, after mice were anaesthetized with isoflurane and decapitated, brains were quickly removed and placed in ice-cold low-sodium, high-sucrose dissecting solution (228 mM sucrose, 26 mM NaHCO 3 , 11 mM glucose, 2.5 mM KCl, 1 mM NaH 2 PO 4 , 1 mM MgSO 4 , 0.5 mM CaCl 2 ). Slices were collected with a Leica VT 1200s vibrating microtome. Slices were allowed to recover for a minimum of 60 min in a submerged holding chamber (∼25 °C) containing artificial cerebrospinal fluid (ACSF) consisting of 119 mM NaCl, 2.5 mM KCl, 2.5 mM CaCl 2 , 1.3 mM MgCl 2 , 1 mM NaH 2 PO 4 , 11 mM glucose and 26.2 mM NaHCO 3 . For hyperplasticity recordings (Extended Data Fig. 6), slices were removed from the holding chamber and placed into the recording chamber, where they were continuously perfused with oxygenated (95% O 2 , 5% CO 2 ) ACSF at 2 ml min−1 at 25 °C. For metaplasticity recordings (Fig. 4), slices were removed from the holding chamber and incubated first for 10 min in oxygenated ACSF containing picrotoxin (50 µM, Sigma), followed by 10-min incubation in oxygenated ACSF containing both picrotoxin and oxytocin (1 µM, Tocris) before being placed into the recording chamber. Whole-cell voltage-clamp recordings from MSNs or layer V pyramidal cells were obtained under visual control using a 40× objective. The NAc core was identified by the presence of the anterior commissure, and the PL/IL region of the mPFC was identified by the presence of the forceps minor of the corpus callosum. Recordings were made with electrodes (2.5–4.0 MΩ) filled with 115 mM CsMeSO 4 , 20 mM CsCl, 10 mM HEPES, 0.6 mM EGTA, 2.5 mM MgCl, 10 mM sodium phosphocreatine, 4 mM sodium ATP, 0.3 mM sodium GTP and 1 mM QX-314. Miniature EPSCs were collected at a holding potential of −70 mV in the presence of tetrodotoxin (0.5 μM, Tocris Biosciences) and picrotoxin (50 μM, Sigma). Two minutes after break-in, 30-s blocks of events (total of 200 events per cell) were acquired and analysed using the Recording Artist plugin in Igor Pro software with threshold parameters set at 5 pA amplitude and <3 ms rise time. All events included in the final data analysis were verified visually. Data were analysed by multivariate analysis of variance (MANOVA) with three independent variables (drug, brain area and age) and two dependent variables (frequency and amplitude). Likelihood ratio test performed comparing the full model using treatment, age, and structure to a reduced model using age and structure. All calculations were performed in either GraphPad Prism 9 or the R programming language and are available as Supplementary Code 1 and in the repository at https://github.com/genesofeve/DolenPsychedelicOpenState.
RNA extraction and sequencing
Male wild-type C57BL6/J mice were injected i.p. with LSD, ketamine, cocaine (20 mg kg−1) or saline solution either 2 weeks or 48 h before the mice were euthanized. At P98 to P112, mice were euthanized, brains were rapidly removed and ~1mm thick coronal slice (n = 3 mice per condition) containing the nucleus accumbens were sectioned using a mouse brain matrix. To microdissect the NAc, slices were placed in a petri dish containing ice-cold ACSF (125 mM NaCl, 2.5 mM KCl, 2 mM CaCl 2 , 1 mM MgCl 2 , 1.25 mM NaH 2 PO 4 , 10 mM glucose and 26 mM NaHCO 3 ) supplemented with RNase inhibitor and oxygenated with carbogen gas (95% O 2 and 5% CO 2 ) to pH 7.3–7.4. The NAc was identified using the anterior commissure and other structural markers. Between each dissection, blades were replaced and all the instruments and the matrix were cleaned with a solution containing RNase inhibitor. Following dissection, tissue was immediately placed into 0.5 ml Trizol and subjected to a 15 s burst with a tissue homogenizer to lyse the cells. Samples were kept on ice prior to storage at −20 °C. Total RNA were extracted using the RNeasy Kit from Qiagen. The quality of purified RNA was assessed via both a nanodrop and 2100 Bioanalyzer from Agilent. Library preparation was performed using a TruSeq Stranded mRNA kit (Illumina) using the recommended protocol. Individual dual-indexed libraries were quality controlled, pooled, and sequenced on the NovaSeq 6000 platform on a single S1 flowcell to an average depth of 76,841,745 (±8,066,939.82) paired-end 100 bp reads per sample. Reads were pseudoaligned to the mouse GENCODE vM25 (ref. 71) reference transcriptome using kallisto (v0.46.2) with 100 bootstrapped samples and 6 threads. Defaults were used for all other parameters. Estimated transcript-level abundances were collapsed to gene-level expression estimates and analysed using the sleuth (v0.30.0) R/Bioconductor package. To identify genes with differential expression as a function of samples where the critical period is reopened we performed a likelihood ratio test comparing a full model which included batch, and critical period to a reduced model that only included batch. Time was not used as an explanatory variable in this model fitting. Using this test, we identified 65 genes as significantly differentially expressed at a 10% false discovery rate (Benjamini–Hochberg-corrected q ≤ 0.1). To identify genes with differential expression as a function of any drug treatment (including cocaine) versus saline we performed a likelihood ratio test comparing a full model that included batch, and ‘treated vs untreated’ to a reduced model that only included batch. Using this test, we identified 39 genes as significantly differentially expressed at a 15% false discovery rate (Benjamini–Hochberg-corrected q ≤ 0.15). Time was not used as an explanatory variable in this model fitting. Raw data will be made publicly available (Gene Expression Omnibus accession numbers: GSE230679 and GSM7231202–GSM7231228). Code to reproduce the RNA-seq analysis and associated figures is provided as Supplementary Code 2 and in the repository at https://github.com/genesofeve/DolenPsychedelicOpenState.
Statistics
All statistical details can be found in the figure legends, including the type of statistical analysis used, P values, n, degrees of freedom, t values and f values. Sample sizes were not predetermined by statistical methods; instead they were estimated based on the previously published literature11. Data distributions were assumed to be normal. Homogeneity of variance was tested using Levene’s test for equality of variances. Comparisons between experimental manipulations were made using a two-tailed Students t-test (paired or unpaired, and with or without Welch’s correction as appropriate) and MANOVA for comparisons between multiple outcome measures, with P < 0.05 considered significant.
Linear, β-spline, loess smoothing and natural spline models evaluated on the previously published time course of normalized social preference scores11. Loess smoothing yielded a pseudoinverse at age 41.695 and a knot point of 35 was chosen for both β-spline and natural spline models. The natural spline outperformed the β-spline (adjusted R2 of 0.1053 versus 0.5554, respectively) with fewer parameters. Residuals were plotted against fitted values and age to check model assumptions. Leave one out cross validation was also used to assess model fit. Control data from all new experiments was used as test data via the predict R function. RSME and R2 values were comparable between the original model and the new data. Two-way t-tests to compare means of controls groups against matched or binned time periods was done to confirm fit to new data. The full model including coefficients for splines, experiment and condition was constructed and tested against reduced models with the final reduced model being reported. MANOVA analysis was carried out using multivariate linear models and the ANOVA function. All statistical comparisons were carried out in the R programming language and can be found in Supplementary Codes 3 and 4 as well as in the repository at https://github.com/genesofeve/DolenPsychedelicOpenState.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.