Appendectomy increases the cumulative incidence of subsequent CRC development of a population-based longitudinal cohort study
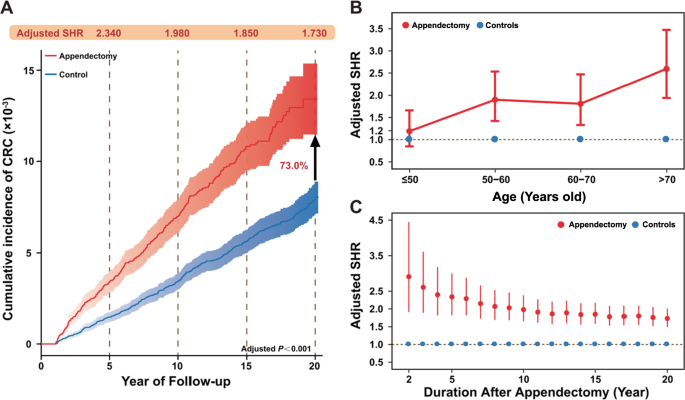
We first conducted a large epidemiological study to explore the association between appendectomy and subsequent CRC risk. A total of 43,976 appendectomy cases and 85,179 age- and gender- non-appendectomy controls were included in further analysis (Fig. S1A). Basic characteristics of these participates were shown in Table S1. During a total of 1,401,020 person-years follow-up time, the CRC incidence was 73.1 (95% CI: 65.0–81.2) per 100,000 person-years in appendectomy group, whereas CRC incidence in controls was 39.7 (95% CI: 35.8–43.7) per 100,000 person-years (Table S2). The overall risk of subsequent CRC development increased by 73.0% in appendectomy cases with sub-distribution hazard ratio (SHR) 1.730 (95% confidence interval (CI) 1.490–2.010) compared with controls (Fig. 1A and Table S2). Subgroup analysis further indicated that appendectomy-treated subjects with aged > 50 years had significantly higher risk for CRC development (SHR 2.020 and 95% CI 1.710–2.396), as compared to appendectomy cases with aged ≤50 years (SHR 1.190 and 95% CI: 0.848–1.660) (Fig. 1B, Table S2 and Fig. S2). In addition, appendectomy cases had significantly higher risk for CRC development in proximal colon (SHR 2.210 and 95% CI: 1.640–2.990) compared with distal colon (SHR 1.670 and 95% CI 1.290–2.150) and rectum (SHR 1.570 and 95% CI 1.210–2.102), respectively (Table S2 and Fig. S2). During the entire follow-up period, the adjusted SHR for CRC development decreased over time, while the risk of CRC in appendectomy cases remained significantly higher than control individuals (Fig. 1C). Subgroup analyses showed the consistent results (Fig. S3).
Fig. 1: The appendectomy increased the overall cumulative risk of colorectal cancer development based on the longitudinal epidemiological study on Hong Kong Cohort. A The cumulative incidence of colorectal cancer (CRC) was increased by 73% in appendectomy cases compared with controls during the 20 years follow-up. B The adjusted sub-distribution hazard ratio (SHR) for CRC development stratified by age (≤50 years old, 50–60 years old, 60–70 years old, and >70 years old). C The temporal trends of the adjusted SHR for CRC development over the 20 follow-up years in appendectomy cases compared with controls. Full size image
Compositional shift of gut microbiome in appendectomy cases
Given the positive clinical association between appendectomy and CRC development, we next investigated the potential contribution of gut dysbiosis induced by appendectomy to the risk of CRC. Gut microbiome profiling was performed in 314 fecal samples from 157 appendectomy cases and 157 normal controls by shotgun metagenomic sequencing (Fig. S1B). Demographic and clinical characteristics of these participants were shown in Table S3. High-quality sequencing reads (mean 15 GB per sample) with an average of 34,168,657 paired reads per sample were obtained after filtering for microbial taxonomic classification (Table S4). Significantly shift in microbial composition was observed at phylum level with relative abundance ≥ 1% (Fig. 2A). In particular, Fusobacteria phylum was markedly enriched in the appendectomy group (Fig. 2B). Subjects with appendectomy had a lower microbial alpha diversity (Simpson index of genera with relative abundance >1%) in their gut microbiome compared to normal controls (P < 0.05) (Fig. 2C). Beta diversity analysis showed separated clusters between appendectomy cases and normal controls (PERMANOVA, P < 0.05; Fig. 2D). To assess whether the microbial signature is influenced by the sampling time, we examined the microbiome in specimens collected 6 to 24 months after appendectomy. The alpha and beta diversities were similar among specimens collected at different timepoints after appendectomy (6 months, 6–12 months, 12–18 months, 18–24 months, and 24 months after appendectomy) (Fig. S4A, B), suggesting that the microbial community changes caused by appendectomy could be persistent for over 2 years. Considering the impact of age on gut microbiome, β-diversity analysis on older (> 50 years) and younger subgroups (≤50 years) was conducted, respectively. Older subjects showed significant difference in microbiome composition between appendectomy cases and controls (PERMANOVA, P = 0.004), while no significant difference was observed among younger subjects (Fig. 2D). These observations suggested that appendectomy leads to alteration in gut microbiome composition, especially for elder individuals.
Fig. 2: The appendectomy altered the microbial community profile. A The microbial composition at phylum level in appendectomy and control groups. Phylum with relative population above 1% was denoted as abundant, while “Other” was denoted as the sum of rare phyla with relative population less than 1%. B Rare phylum composition in appendectomy and control subjects. “Other” represented the sum of phyla with relative population less than 0.01%. C Alpha diversity for Simpson index at genus level. D Principal coordinates analysis for all subjects, young subjects (age ≤ 50), and old subjects (age > 50). Red stands for the appendectomy group, while blue represents the control group. Full size image
Appendectomy induces enrichments of CRC-associated species in gut
We then extended our microbial analysis by evaluating the difference in gut microbiome at species level. 25 bacterial species were identified with significant difference in abundances (11 enriched and 14 depleted) in appendectomy cases compared with control subjects (Fig. 3A). Among the 11 enriched species, 7 were reported as CRC- or cancers-associated bacteria (Bacteroides vulgatus, Bacteroides fragilis, Villanelle dispar, Prevotella ruminicola, Prevotella fusca, Prevotella dentalis, and Prevotella denticola) [6, 13,14,15,16,17,18]. While 5 were reported as protective bacteria (Blautia sp SC05B48, Collinsella aerofaciens, Lachnospiraceae bacterium Choco86, Enterococcus hirae, Blautia sp YL58) [15, 19,20,21,22] among the 14 depleted species (Fig. 3A and Figs. S5, S6). In particular, the top two enriched bacteria in appendectomy subjects, B. vulgatus and B. fragilis, were found to have age-specific correlation, in which they were significantly enriched only in the older subgroup (both P < 0.001) (Fig. 3B). The top two depleted bacteria in appendectomy, B. sp SC05B48 and C. aerofaciens, also showed age-specific correlation with significant depletion in the younger subgroup (both P < 0.01) (Fig. 3B). Such changes in bacterial abundances (B. vulgatus, B. fragilis, B. sp SC05B48 and C. aerofaciens) were sustained for over 2 years after appendectomy (Fig. S7A). Moreover, the virulence factor gene pks was significantly increased (P < 0.05) in appendectomy cohort compared to normal controls (Fig. S7B). To further validate the abundance of differentially enriched bacteria identified by our metagenomic analyses, we performed qPCR of 3 representative enriched (B.fragilis, B. vulgatus, V. dispar) and depleted bacteria (E. hirae, Lachnospiraceae bacterium Choco86, Blautia sp. SC05B48) in appendectomy cases. The results showed a consistent distribution of differential enriched bacterial species as metagenomic analyses (Fig. 3C and Fig. S8). Collectively, these observations consistently indicated that alterations in gut microbiome after appendectomy could be pathogenic with enrichments of several CRC-associated bacteria and depletion of beneficial commensals.
Fig. 3: Differential abundance of bacterium species in appendectomy patients compared to control subjects. A Bar plot for the differential abundant species identified by LEfSe. Red bars stand for the enriched species in appendectomy patients, while blue bars represent the depleted species in appendectomy subjects. Species that were reported to promote cancer were marked in bold red fonts, while the cancer-inhibiting species were marked in bold green fonts. B The relative abundance for representative appendectomy enriched species, Bacteroides vulgatus and Bacteroides fragilis; and appendectomy-depleted species, Blautia sp SC05B48 and Collinsella aerofaciens in all subjects, young and old subgroups are compared and shown as box plot. C The validation of identified appendectomy-enriched and depleted bacteria species including Bacteroides fragilis, Bacteroides vulgatus, Veillonella dispar, Enterococcus hirae, Lachnospiraceae bacterium Choco86, Blautia sp. SC05B48) was shown by using targeted qPCR in appendectomy cases and control subjects. Full size image
Microbiome ecological network is altered after appendectomy
To gain insights into ecological networks among bacteria, we investigated correlations between enriched and depleted species in appendectomy subjects by Spearman’s Rank correlation analyses. We found that enriched and depleted bacteria separately formed their own networks with negative correlation with each other (Fig. 4A, B). While the number of negative correlations was significantly increased in the appendectomy group at the ratio of 20:4 ratio, contrasting to the ratio of 2:9 for positive correlations (Fig. 4A, B, Tables S5,S6). In particular, we identified that B. fragilis and B. vulgatus, two of appendectomy-enriched species and the CRC-associated pathogen [14, 16], was the centrality of the network of enriched bacteria and had the strongest negative correlation with the network of depleted bacteria (Fig. 4A, C, Table S7). Whereas appendectomy-depleted bacteria showed significant synergistic relationships (Fig. 4A, C), possibly due to the protective effects of beneficial bacteria in the gut microenvironment. Meanwhile, in consistent with the epidemiological results, we observed more negative correlations among appendectomy-enriched and -depleted species in older subjects than younger subjects (Fig. 4C). The appendectomy-enriched bacteria, including B. fragilis and B. vulgatus, showed increased negative correlations with commensal bacteria in the older subgroup, while protective bacteria such as Blautia sp. SC05B45 and Blautia sp. YL58 had negligible influence. Moreover, correlations among appendectomy-depleted bacteria were more positive in the younger subgroup than in the older subgroup. Taken together, the change in gut microbial ecology after appendectomy indicated that appendectomy could impact interactions among gut microbes which may contribute to CRC development in appendectomy cases, especially in aged subjects.
Fig. 4: Correlation analysis of differential abundant bacteria species. A Correlation network among differential abundant species (enriched/depleted) selected by LEfSe in appendectomy and control groups. B Heatmap for all stronger relationships in appendectomy cases among appendectomy-enriched/depleted bacteria. The node size in the heatmap represented Spearman’s rank correlation coefficient. C Subgroup analysis (age ≤50/ age > 50 years) of associations in appendectomy cases among selected bacteria with correlation coefficient rho below −0.2 or above 0.6 are shown. Correlations were calculated by Spearman’s rank correlation. The node size in the networks represents log2 (Fold Change). The red lines represent co-exclusion correlation, and green lines represent co-occurrence association. Species that were reported to promote cancer were marked in red fonts, while the cancer-inhibiting species were marked in blue fonts. Full size image
Functional features of gut microbiome are altered after appendectomy
The contribution of gut microbial dysbiosis to pathological conditions could also be mediated by their functional capabilities. Functional changes at species-level were characterized by Kyoto Encyclopedia of Genes and Genomes (KEGG) database, and a total of 227 KEGG orthology genes (KO genes) showed significant enrichments in appendectomy group, compared to healthy controls (adjusted P < 0.1). We then examined abundances of microbial genes to assess the functional roles of gut microbes in appendectomy-associated CRC risk. Functional pathways including amine acid metabolism, carbon fixation, sulfur metabolism, and aromatic amino acid metabolism were significantly different in appendectomy cases (Fig. 5A), of which all of them were reported to be associated with CRC development [23,24,25,26]. Changes in the functional microbial pathway were examined using HUMAnN2, which enables robust profiling on microbial pathway abundance in distinct metagenome functional contents. We identified 15 metabolic pathways with differential abundances between appendectomy cases and controls (q < 0.2). Microbiome in appendectomy cases was dominated by biosynthesis pathways of deoxyribonucleotides (pyrimidine, adenosine, guanosine), peptidoglycan, L-glutamate L-glutamine, and pyrimidine deoxyribonucleotides (P < 0.01, for all), which were all reported as cancer-promoting metabolic pathways [27,28,29]. Moreover, the biosynthesis of L-proline from arginine was known to be cancer-inhibitory and was found to be depleted in appendectomy cases [26] (P < 0.0001) (Fig. 5B).
Fig. 5: Microbiome functional capacity changes in appendectomy patients. A Appendectomy-induced alteration of abundance in microbial genes was summarized by KEGG pathway modules. The genes enriched in appendectomy compared to normal control were evaluated for gene relative abundance profiles (Wilcoxon rank-sum test followed by FDR correction using “Benjamini–Hochberg” methods, q < 0.1), and those with mean relative abundance >1% were shown. The relative gene abundance was shown as bar plots by averaging all appendectomy samples (n = 157). B Bar plot for Pathway Enrichment Analysis by Humann2 with Meta-Cyc database. The red bar standard for appendectomy-enriched pathways compared with control group, whist the blue bar represented the appendectomy-depleted pathways. Pathways reported to promote cancer were marked in red fonts, and the cancer-inhibiting ones were marked in green fonts. Full size image
Appendectomy promotes colorectal tumorigenesis through altering microbial composition and inducing intestinal barrier dysfunction in mice
To confirm the influence of appendectomy on microbial dysbiosis and colorectal tumorigenesis, we performed appendectomy or sham in a carcinogen-induced CRC mouse model (Fig. 6A). The results showed that tumor number (P < 0.05) and tumor size (P < 0.05) were significantly higher in both male and female mice with appendectomy as compared to control mice (Fig. 6B), while no significant changes in body weight was observed (Fig. S9A). Histological examination of colon tumor sections confirmed that appendectomy induced cell proliferation as evidenced by significantly increased Ki-67 positive cells (Fig. 6C).
Fig. 6: Appendectomy promotes colorectal tumorigenesis through altering microbial composition and inducing intestinal barrier dysfunction in mice. A Schematic diagram of mice experiment. Appendectomy and control were performed in a AOM/DSS-induced CRC mouse model. B Representative images of colorectal tumors from mice with control or appendectomy (top) and H&E staining of tumors (bottom). The tumor load was evaluated by tumor number and tumor size (mm3). C Immunohistochemistry staining of Ki-67+ cells in colon tumors of appendectomy or control mice. D Gut microbial alpha diversity by richness and Chao1 and the beta-diversity measured by dissimilarity using bray-cutis distance in stool samples collected at baseline (3 days before appendectomy), and 3, 7, 14, 30 days after appendectomy. E Differentially enriched gut bacteria before and after appendectomy in mice. F Serum lipopolysaccharides concentration and Western blot analysis of Zo-1, Occludin, and E-cadherin in colon of Appendectomy and control mice. Band intensity was measured by Image J and the ratio of each band was normalized to the corresponding GAPDH. G The E-cadherin protein expression in colon tumor from appendectomy and control mice. Full size image
Fecal samples of mice at different time points including baseline (3 days before appendectomy), and 3, 7, 14, 30 days after appendectomy, were subjected to metagenome sequencing for investigating the effects of appendectomy on gut microbiome. Compared to baseline, the microbial alpha diversity (Richness and Chao1) was significantly decreased 3 days after appendectomy (Fig. 6D). Significant difference in beta-diversity as measured by dissimilarity using bray-cutis distance was also observed between baseline and 3 days after appendectomy (Fig. 6D). In contrast, the microbial diversity (alpha and beta-diversity) was similar among fecal samples collected 3, 7, 14 and 30 days after appendectomy (Fig. 6D), suggesting the microbiome changes could persist after appendectomy. Moreover, some pathogenic bacteria, including Bacteroides intestinalis and Shigella sonnei, were enriched after appendectomy (Fig. 6E and Table S8).
We further examined the intestinal barrier function, which is known to be greatly impaired in colorectal tumorigenesis, in mice receiving appendectomy. Serum lipopolysaccharides concentration was significantly elevated in mice with appendectomy compared to control mice (Fig. 6F). In keeping with this, the expressions of tight junction proteins Zo-1 (P < 0.01), Occludin (P < 0.001), and adherent junction protein E-cadherin (P < 0.001) were all significantly reduced in the colon of appendectomy mice (Fig. 6F), implying a remarkable impairment of intestinal barrier function in appendectomy mice. Moreover, the protein expression of E-cadherin was decreased in tumor tissues of post-appendectomy mice compared to control (P = 0.01, Fig. 6G). Accordingly, the E-cadherin inhibited oncogenic Hippo signaling [30], was activated in tumor tissue of post-appendectomy mice compared to the control mice as identified by RNA sequencing (Fig. S9B). Collectively, these results indicated the CRC-promoting role of appendectomy was associated with gut dysbiosis through impairing intestinal barrier function and downregulated E-cadherin signaling.
To verify the roles of gut bacteria in promoting CRC after appendectomy, we depleted the gut microbiome by antibiotic cocktail treatment in both male and female mice (Fig. S10A). The depletion of gut microbiome by antibiotics was confirmed by qPCR on mouse fecal samples (Fig. S10B). No difference in body weight was found between control and appendectomy groups with antibiotic treatment (Fig. S10C). Appendectomy could not affect CRC development in microbiome-depleted mice, which had similar levels of tumor number and tumor size as in antibiotic-treated mice with sham, regardless of their gender (Fig. S10D). These findings corroborated that intestinal bacteria dysbiosis caused by appendectomy play a key role in CRC tumorigenesis after appendectomy.